Welcome to Charu G. Kumar's Group
Welcome to our group's website which highlights our research focus in the areas of Systems and Computational Biology. We are part of the Department of Bioengineering, University of Illinois at Urbana-Champaign. Research in our group is highly interdisciplinary using computational genomics, bioinformatics, data mining, evolutionary, and systems biology approaches. The major thrust areas of our research are:
Microbiome/Pathogen Computational Genomics and metabolic network modeling
Many diseases are a result of environmental causes that are sourced at sites distal to the disease target. For example, infection may initiate in the soil/water as a result of bacterial degradation of an animal product. These bacteria may be dispersed via spores, or they may be transferred via host animal/insects to human targets, resulting in disease. We study changes in metabolism of these pathogenic bacteria/microbiome as model organism/communities and as a function of environmental catalysts to disease. Of particlar interest is to identify differentially correlated pathways in gene metabolic networks from environmental microbiome samples. The goal is to identify relationships between environmental stresses, microbial biogeography and human zoonotic diseases.Complex neurological diseases
We use computational neuroscience and systems biology approaches to study complex diseases that are known to differentially affect astrocytes. At present we are focused on glioblastoma, fragile X and autism. The goal is to develop an understanding of the role played by astrocytes in relation to the function (and dysfunction) of the entire brain. Questions of interest include studying how these diseases may have created molecular novelties operant in the constituent pathways and what mechanisms are at play in generating these novelties, which could then be studied for their potential as drug-targets.Computational evolutionary biology
A third research direction in our group is studying the origin and evolution of metabolic, regulatory, and signaling networks and modules at a genome wide level in humans, in model organisms, and in pathogenic bacteria. We integrate comparative genomics, '-omics' data mining, and systems biology approaches to explore comparative evolution of gene networks between organisms.
Charu G. Kumar
Systems and Computational Biology Group
Department of Bioengineering
Faculty affiliate, Neuroscience program
University of Illinois at Urbana-Champaign
3146D Everitt Lab
1406 W. Green St.
Urbana, IL 61801
E-mail: cgkumar at illinois.edu
Updated:
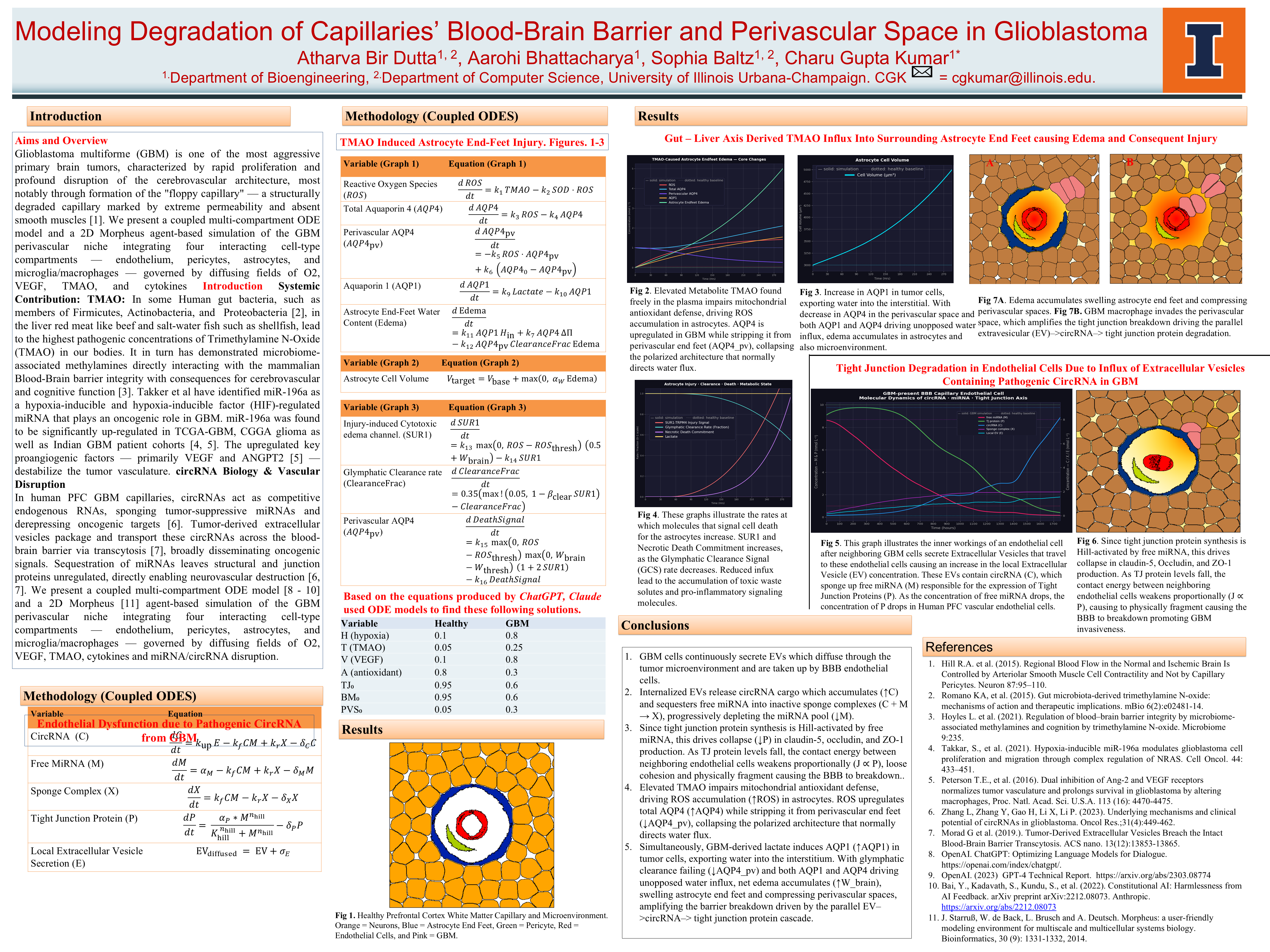
New POSTER for the Undergraduate Research Symposium in the Personal page:
These are two PDFs on Microbes, Diseases, and nutrition that you may download:
Microbes_Diseases_First_06062024.pdf
Microbes_Nutrition_Second_2282020.pdf
Updated:
New POSTER for the Undergraduate Research Symposium: